Add training option unique_pairs#268
Conversation
Co-authored-by: Tom Aarsen <37621491+tomaarsen@users.noreply.github.com>
Co-authored-by: Tom Aarsen <37621491+tomaarsen@users.noreply.github.com>
Co-authored-by: Tom Aarsen <37621491+tomaarsen@users.noreply.github.com>
|
|
First of all, these changes look great! I way prefer simply having train_examples = sentence_pairs_generation(...)rather than train_examples = []
for _ in range(...):
train_examples = sentence_pairs_generation(..., train_examples)I ran some quick experiments locally, and I noticed an interesting quirk. Before I stress you out: this quirk also exists on main. If I increase So, whereas I might get 34 training steps times 1 epoch (so, 34 steps total) with Simple script to reproduce the quirkfrom datasets import load_dataset
from setfit import SetFitModel, SetFitTrainer, sample_dataset
# Load a dataset from the Hugging Face Hub
dataset = load_dataset("sst2")
# Simulate the few-shot regime by sampling 8 examples per class
train_dataset = sample_dataset(dataset["train"], label_column="label", num_samples=8)
eval_dataset = dataset["validation"]
# Load a SetFit model from Hub
model: SetFitModel = SetFitModel.from_pretrained(
"sentence-transformers/paraphrase-mpnet-base-v2",
)
# Create trainer
trainer = SetFitTrainer(
model=model,
train_dataset=train_dataset,
eval_dataset=eval_dataset,
column_mapping={"sentence": "text", "label": "label"}, # Map dataset columns to text/label expected by trainer
num_epochs=1, # <- Change 1 to see the quirk
)
# Train and evaluate
trainer.train()
metrics = trainer.evaluate()It's caused by this line: Line 378 in f777c2c The train_steps is defined like so:Line 366 in f777c2c Which explains the behaviour. In my opinion, the number of steps per epoch should be equivalent to the number of datapoints in the dataloader. That is exactly the behaviour if steps_per_epoch is not supplied. So, could you remove supplying the steps_per_epoch parameter in your PR? That way, we can prevent this quirk.
Experiments on sst2As mentioned, I ran some experiments. In particular, I used unique_pairs = False w. 3 epochs of 640 steps (40 batches) each (1920 total optimization steps) (Baseline)from datasets import load_dataset
from setfit import SetFitModel, SetFitTrainer, sample_dataset
# Load a dataset from the Hugging Face Hub
dataset = load_dataset("sst2")
# Simulate the few-shot regime by sampling 8 examples per class
train_dataset = sample_dataset(dataset["train"], label_column="label", num_samples=8)
eval_dataset = dataset["validation"]
# Load a SetFit model from Hub
model: SetFitModel = SetFitModel.from_pretrained(
"sentence-transformers/paraphrase-mpnet-base-v2",
)
# Create trainer
trainer = SetFitTrainer(
model=model,
train_dataset=train_dataset,
eval_dataset=eval_dataset,
column_mapping={"sentence": "text", "label": "label"}, # Map dataset columns to text/label expected by trainer
num_epochs=3,
unique_pairs=False,
seed=1,
)
# Train and evaluate
trainer.train()
metrics = trainer.evaluate()unique_pairs = True w. 14 epochs of 136 steps (9 batches) each (1904 total optimization steps)from datasets import load_dataset
from setfit import SetFitModel, SetFitTrainer, sample_dataset
# Load a dataset from the Hugging Face Hub
dataset = load_dataset("sst2")
# Simulate the few-shot regime by sampling 8 examples per class
train_dataset = sample_dataset(dataset["train"], label_column="label", num_samples=8)
eval_dataset = dataset["validation"]
# Load a SetFit model from Hub
model: SetFitModel = SetFitModel.from_pretrained(
"sentence-transformers/paraphrase-mpnet-base-v2",
)
# Create trainer
trainer = SetFitTrainer(
model=model,
train_dataset=train_dataset,
eval_dataset=eval_dataset,
column_mapping={"sentence": "text", "label": "label"}, # Map dataset columns to text/label expected by trainer
num_epochs=14,
unique_pairs=True,
seed=1,
)
# Train and evaluate
trainer.train()
metrics = trainer.evaluate()Results
Notes:
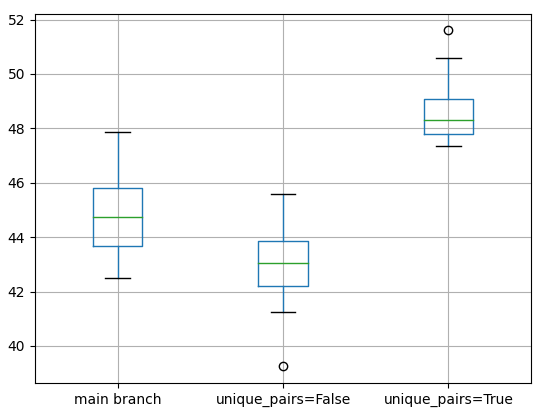
The results are almost statistically significant at a p-value of .074996 (assuming p < .05 for statistical significance), but certainly very promising. Especially the difference in standard deviation is very promising for this PR, as it displays the expected behaviour that non-random sampling produces more stable and consistent results. I'm quite excited about it. I'm considering running more experiments. Feel free to run your own ones, too!
|
Experiments on emotionI ran some more experiments. In particular, I used These experiments differ from the previous experiments on sst2 in that each epoch ran the same number of examples, and that I used the same number of epochs between the experiments. Furthermore, sst2 is binary, while the emotion dataset contains 6 classes. unique_pairs = False w. num_iterations=10 and 2 epochs of 960 steps (60 batches) each (1920 total optimization steps) (Baseline)from datasets import load_dataset
from setfit import SetFitModel, SetFitTrainer, sample_dataset
# Load a dataset from the Hugging Face Hub
dataset = load_dataset("SetFit/emotion")
# Simulate the few-shot regime by sampling 8 examples per class
train_dataset = sample_dataset(dataset["train"], label_column="label", num_samples=8)
eval_dataset = dataset["validation"]
for seed in range(12):
# Load a SetFit model from Hub
model: SetFitModel = SetFitModel.from_pretrained(
"sentence-transformers/paraphrase-mpnet-base-v2",
)
# Create trainer
trainer = SetFitTrainer(
model=model,
train_dataset=train_dataset,
eval_dataset=eval_dataset,
num_epochs=2,
num_iterations=10,
unique_pairs=False,
seed=seed
)
# Train and evaluate
trainer.train()
metrics = trainer.evaluate()
print(metrics)unique_pairs = True w. num_iterations=10 and 2 epochs of 960 steps (60 batches) each (1920 total optimization steps)from datasets import load_dataset
from setfit import SetFitModel, SetFitTrainer, sample_dataset
# Load a dataset from the Hugging Face Hub
dataset = load_dataset("SetFit/emotion")
# Simulate the few-shot regime by sampling 8 examples per class
train_dataset = sample_dataset(dataset["train"], label_column="label", num_samples=8)
eval_dataset = dataset["validation"]
for seed in range(12):
# Load a SetFit model from Hub
model: SetFitModel = SetFitModel.from_pretrained(
"sentence-transformers/paraphrase-mpnet-base-v2",
)
# Create trainer
trainer = SetFitTrainer(
model=model,
train_dataset=train_dataset,
eval_dataset=eval_dataset,
num_epochs=2,
num_iterations=10,
unique_pairs=True,
seed=seed
)
# Train and evaluate
trainer.train()
metrics = trainer.evaluate()
print(metrics)Results
Notes:
These results are very much statistically significant! These seem like serious improvements. And again, we can note an increase in stability as However, from these results it is also clear that the performance of I'm very positive about these results.
|
|
Dan and I had a discussion on this topic, which I will share for others reading through this interesting PR: We theorize that the reduction in accuracy when Additionally, Dan tested equal representation of positive pairs versus negative pairs compared to the Furthermore, I'm very interested in "freshly" generating samples per epoch. For example, if there are 20.000 possible negative pairs, and we sample 2.000 in a 20-epoch training scenario, then currently we use the same 2,000 samples for each of the 20 epochs. We might get an increase in performance if we can re-sample 2.000 from the 20.000 for every epoch again.
|
|
Non-representative sampling fix - Overall the new sampler improves reproduciblity and samples labelled datasets more evenly, with slight accuracy & reducing std dev as a result:
Setting up unique_pairs - forcing use of equal pos/ neg samples shows better results but requires sampling "unique pairs" differently...
Results on the multi-class (imbalanced) test datasets (opt_steps are a proxy for runtime), default 1 epoch:
Results show increasing no. of epochs improves accuracy in roughly similar runtime, but not as effective as more sampling. As noted in the last comment in this PR, generating samples for each epoch run would be nice addition, however epochs are handled by the |
I'm lending towards the latter due to the higher accuracy and provides a natural limit to the no. of samples worth used in training. The |
|
@danstan5 My first hunch is to prefer oversampling rather than undersampling, for two reasons:
That said, As for the merge conflicts, no need to worry about them.
|
|
Nice one. Strong agree on the oversampling.. struggling for good names though ( Sure, I think it's best placed in a more major release as it will ultimately change existing results if your re-train setfit models. Thanks, Dan |
|
Sounds good. We'll have to think of a strong name. Alternatively, we can consider more major changes: Only supporting the oversampling. After all, how much does that really differ from the current behaviour? We could also remove I'm trying to think if there are any scenarios that would be possible now that won't be possible if we take that potential approach, and whether the performance would be degraded in any of the scenarios. Another approach is to take it a step further and take new pairs at the start of every epoch. That said, if we indeed go with the oversampling solution, then either the positive or the negative pairs will simply be all of the unique pairs (assuming I'm curious about your thoughts on this.
|
|
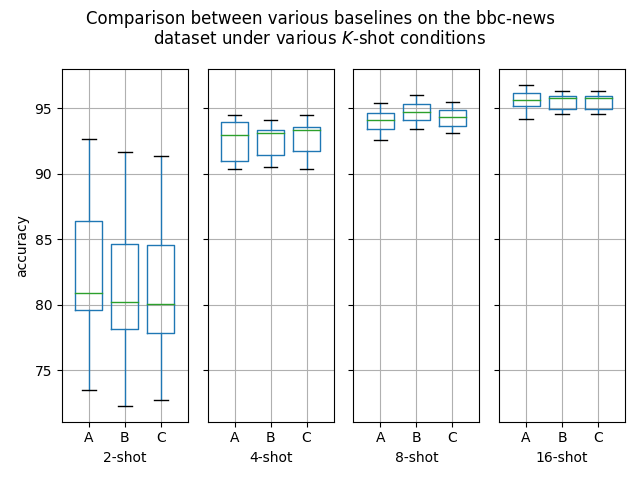
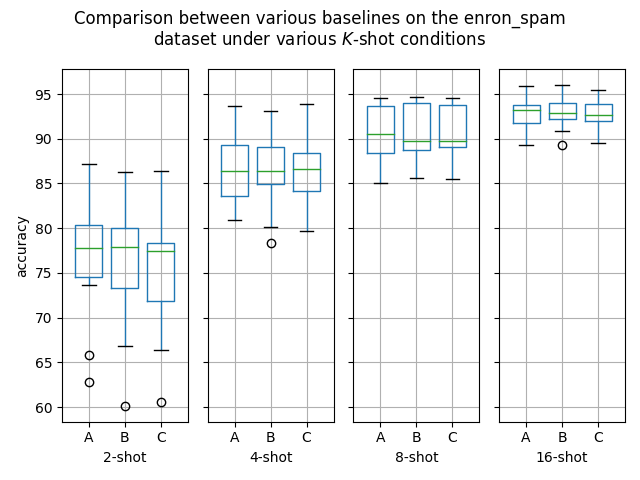
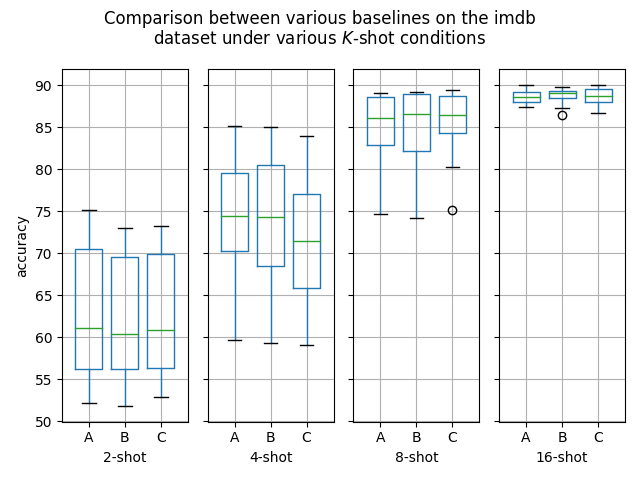
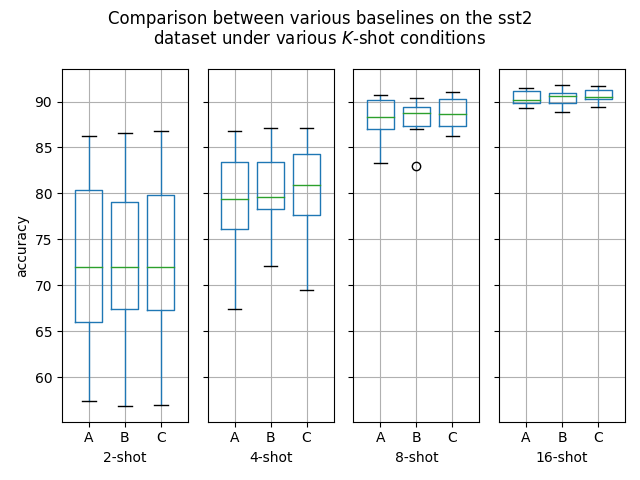
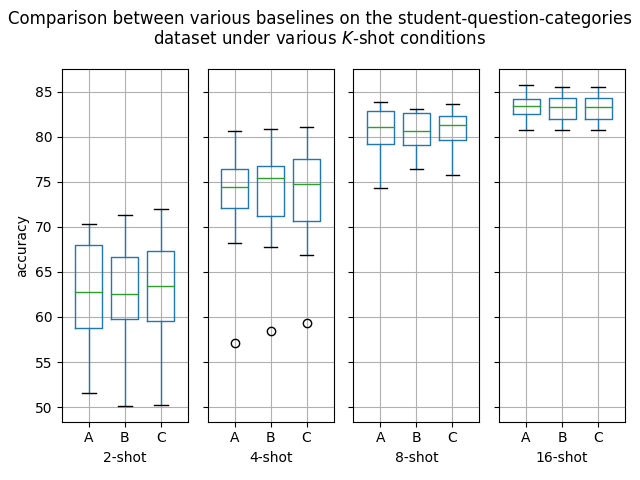
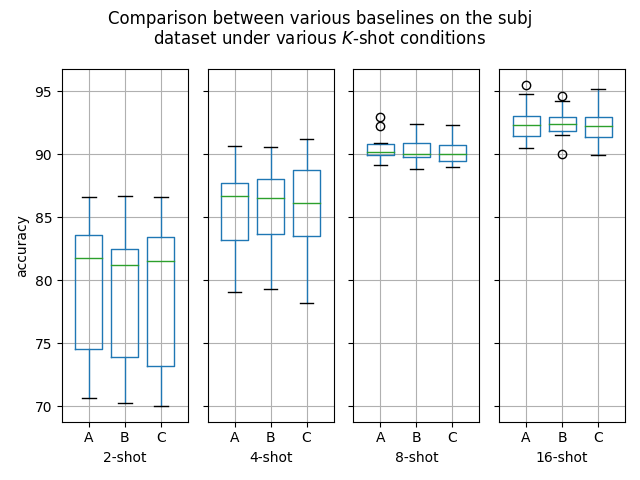
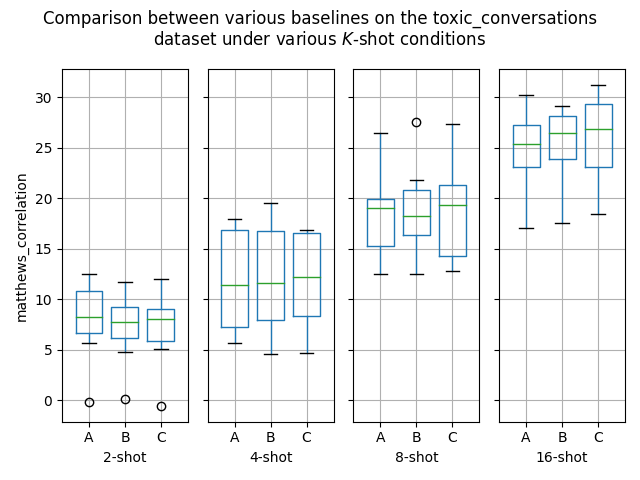
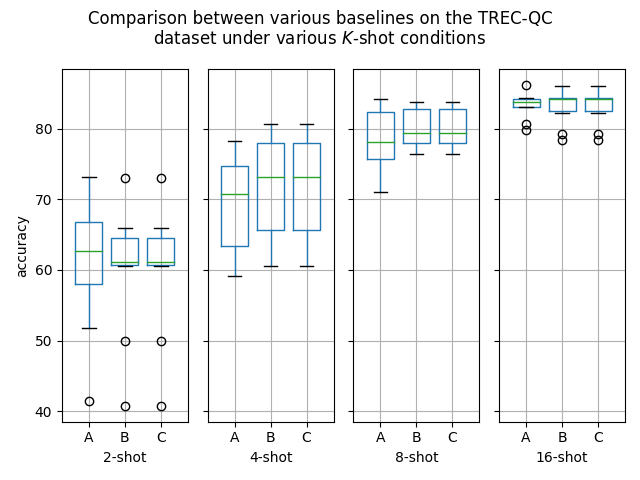
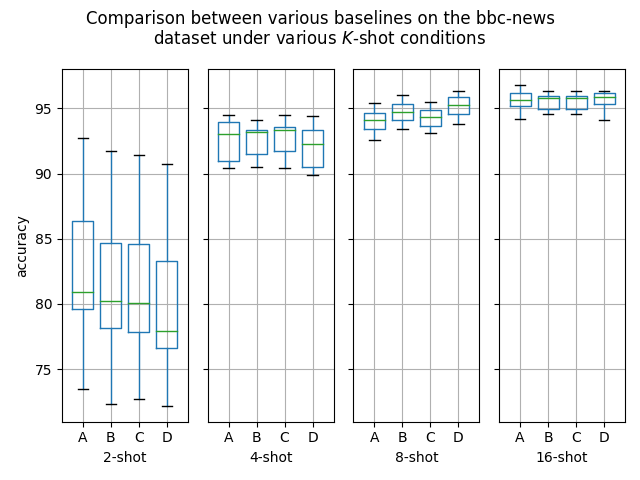
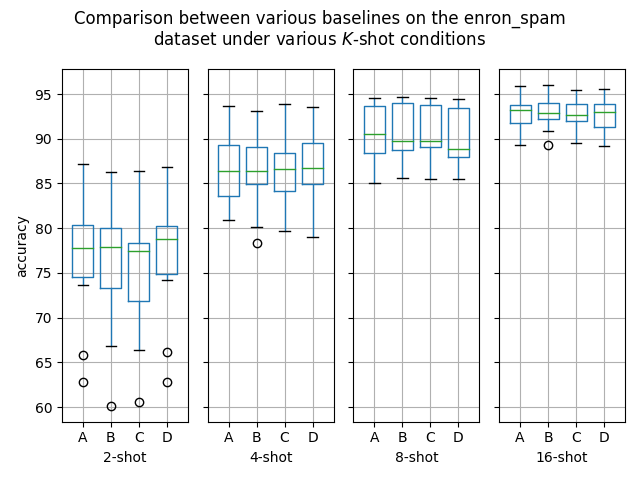
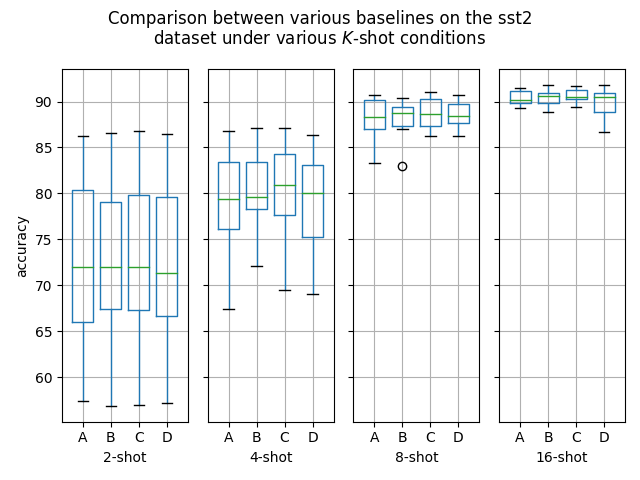
Hello! I've ran some more (much more thorough) experiments on this PR, as myself and others suspected that this PR might help with an ubiquitous overfitting problem on SetFit models. After all, better data gives better models, and conceptually this PR should result in better data. Experiment detailsI have ran three sets of experiments:
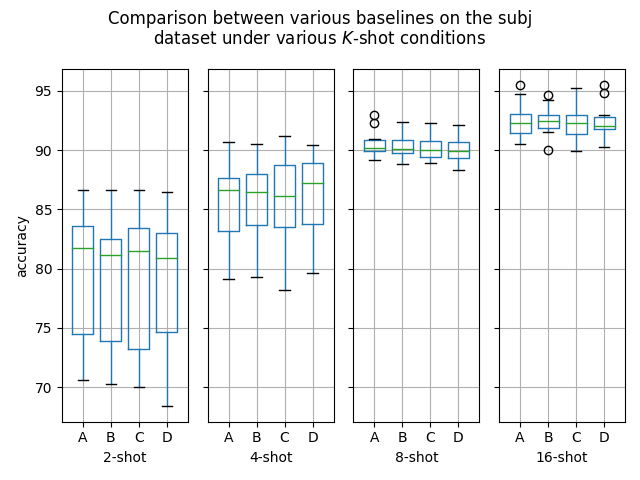
GoalsI have also written a script to plot the results from the three separate ResultsDiscussionI am unable to see significant differences between the three approaches tested here, which is quite a shame. These results are quite surprising to me, as in my previous experiments [1] [2], I encountered improved performance when considering the same number of iterations. I see three reasonable explanations:
I'd like to further investigate option 2 this afternoon. @danstan5 Please let me know if you spot a conceptual flaw in my experiments here. I may have overlooked something. I'd like to notify @danielkorat and @MosheWasserb of these results, as I think they may benefit from the results of this experiment. I also think that @lewtun may be interested in this PR and its various experiments.
|
|
Hey @tomaarsen love these plots, can you share the script for them?? Your right there's nothing to statistically difference in these results, but this is because the default The benefits of this PR only really only come at low |
|
To add on to my previous comment, it does not seem like the performance degraded since earlier in this PR. ExperimentI ran another experiment:
ResultsDiscussionAs before, the performance seems nearly identical to the other results, despite now using the older version of this PR. I'm curious to hear your thoughts,
|
Wonderful! I'll try to see if I can experiment a bit further in those cases. I was under the impression that this PR was most interesting in small dataset situations. Most of all I'm a bit saddened that I can't seem to reproduce my earlier results where this PR seemed to result in a ~3% accuracy jump from 45% to 48% on the emotion dataset. As for the script I used, I'll attach it here now, but I'll also fix up the comments, imports etc. (e.g. some comments talk about Click to see plotting scriptimport argparse
import json
import os
from pathlib import Path
from collections import defaultdict
from glob import glob
from typing import List, Tuple
import matplotlib.pyplot as plt
import pandas as pd
import string
def get_sample_sizes(path: str) -> List[str]:
return sorted(list({int(name.split("-")[-2]) for name in glob(f"{path}/*/train-*-0")}))
def get_formatted_ds_metrics(path: str, dataset: str, sample_sizes: List[str]) -> Tuple[str, List[str]]:
split_metrics = defaultdict(list)
for sample_size in sample_sizes:
result_jsons = sorted(glob(os.path.join(path, dataset, f"train-{sample_size}-*", "results.json")))
for result_json in result_jsons:
with open(result_json) as f:
result_dict = json.load(f)
metric_name = result_dict.get("measure", "N/A")
split_metrics[sample_size].append(result_dict["score"])
return metric_name, split_metrics
def plot_summary_comparison(paths: List[str]) -> None:
dataset_to_df = defaultdict(pd.DataFrame)
dataset_to_metric = {}
for path_index, path in enumerate(paths):
ds_to_metric, this_dataset_to_df = get_summary_df(path)
for dataset, df in this_dataset_to_df.items():
df["path_index"] = path_index
dataset_to_df[dataset] = pd.concat((dataset_to_df[dataset], df))
dataset_to_metric = dataset_to_metric | ds_to_metric
# Prepare folder for storing figures
image_dir = Path("scripts") / "images"
image_dir.mkdir(exist_ok=True)
new_version = max([int(path.name[2:]) for path in image_dir.glob("v_*/") if path.name[2:].isdigit()], default=0) + 1
output_dir = image_dir / f"v_{new_version}"
output_dir.mkdir()
for dataset, df in dataset_to_df.items():
columns = [column for column in df.columns if not column.startswith("path")]
fig, axes = plt.subplots(ncols=len(columns), sharey=True)
for column_index, column in enumerate(columns):
ax = axes[column_index]
# Set the y label only for the first column
if column_index == 0:
ax.set_ylabel(dataset_to_metric[dataset])
# Set positions to 0, 0.25, ..., one position per boxplot
# This places the boxplots closer together
n_boxplots = len(df["path_index"].unique())
allotted_box_width = 0.2
positions = [allotted_box_width * i for i in range(n_boxplots)]
ax.set_xlim(-allotted_box_width * 0.75, allotted_box_width * (n_boxplots - 0.25))
# ax.set_xticks(range(n_boxplots), rotation="vertical")
df[[column, "path_index"]].groupby("path_index", sort=True).boxplot(
subplots=False, ax=ax, column=column, positions=positions
)
k_shot = column.split("-")[-1]
ax.set_xlabel(f"{k_shot}-shot")
if n_boxplots > 1:
# If there are multiple boxplots, override the labels at the bottom generated by pandas
if n_boxplots <= 26:
ax.set_xticklabels(string.ascii_uppercase[:n_boxplots])
else:
ax.set_xticklabels(range(n_boxplots))
else:
# Otherwise, just remove the xticks
ax.tick_params(labelbottom=False)
if n_boxplots > 1:
fig.suptitle(f"Comparison between various baselines on the {dataset}\ndataset under various $K$-shot conditions")
else:
fig.suptitle(f"Results on the {dataset} dataset under various $K$-shot conditions")
fig.tight_layout()
plt.savefig(str(output_dir / dataset))
def get_summary_df(path: str) -> None:
"""Given per-split results, creates a summary table of all datasets,
with average metrics and standard deviations.
Args:
path: path to per-split results: either `scripts/{method_name}/{results}/{model_name}`,
or `final_results/{method_name}/{model_name}.tar.gz`
"""
sample_sizes = get_sample_sizes(path)
header_row = ["dataset", "measure"]
for sample_size in sample_sizes:
header_row.append(f"{sample_size}_avg")
header_row.append(f"{sample_size}_std")
dataset_to_metric = {}
dataset_to_df = {}
for dataset in next(os.walk(path))[1]:
metric_name, split_metrics = get_formatted_ds_metrics(path, dataset, sample_sizes)
dataset_df = pd.DataFrame(split_metrics.values(), index=[f"{dataset}-{key}" for key in split_metrics]).T
dataset_to_metric[dataset] = metric_name
dataset_to_df[dataset] = dataset_df
return dataset_to_metric, dataset_to_df
def main() -> None:
parser = argparse.ArgumentParser()
parser.add_argument("--paths", nargs="+", type=str)
args = parser.parse_args()
plot_summary_comparison(args.paths)
if __name__ == "__main__":
main()When calling this script, I provide it with the multiple paths like so:
|
|
More commits!
These changes are a solution to the above:
Breaking changes:
Impact:
Thoughts as ever appreciated @tomaarsen on this. I can get some of the code a bit clearer, add Distillation sampler into similar class, update tests etc. but keen to get thoughts first |
|
I'm a great fan of how this is looking! I don't have a lot of time to dive into it now (I'll look on Monday), but is my initial assumption correct that with these changes in place, increasing |
As used in comments in huggingface#268
* Add comparison plotting script As used in comments in #268 * Apply automatic formatting * Write the command used to plot the graphs to the output directory
|
I ran some tests with the new
This affords us to deprecate one of them (i.e. I'd love to know your thoughts on this. Edit: I'm unsure about the time efficiency consequences of using a generator, i.e. of having to do a bunch of CPU calls every iteration rather than once in bulk at the start of each epoch.
|
|
The experiments here look very interesting and promising. @tomaarsen - would you please tell me if these changes are going to be merged soon? |
|
This has been superseded by tomaarsen#5, which has been merged into
|
Derived from #259
Changes:
max_pairscount (set bynum_iterations). If all unique pairs have been generated andunique_pairs=Trueit will stop at this point (logging a warning).unique_pairs=Trueneg/ pos pair counts will be remain balanced until all unique positive pairs have been added, then only unique neg pairs will continue to be added. Note: if concerned about an imbalance of negative pairs (in case of high no. of classes)num_iterationscan simply be decreased to balance this as desired.Overall these sampler changes would improve reproducibility + provides more representative sampling of the dataset, useful in cases of strong class imbalance or samples >> no. iterations training.
unique_pairsadds an option for efficient embedding training by maximising only the available data for quicker training (testing for confirmation on this to follow..)Still todo:
test_set- runtime/ accuracy comparisons@tomaarsen keen to get your thoughts on this so far 👍