A new version of the FAST (33 bins) Spectral-Bin Microphysics (FSBM)#848
A new version of the FAST (33 bins) Spectral-Bin Microphysics (FSBM)#848JS-WRF-SBM wants to merge 15 commits intowrf-model:developfrom JS-WRF-SBM:newFSBM
Conversation
|
@JS-WRF-SBM On the github page for this PR, where you see the EDIT box to the right of your title, click on the box that has "wrf-model:release-v4.0.3". Change this to "wrf-model:develop". That is why it says you are changing 535 files, when you want to change about 12. |
|
@JS-WRF-SBM |
|
@JS-WRF-SBM |
|
@JS-WRF-SBM
|
|
Dave, I'm trying to EDIT and than choose from the box the "develop" option, by I'm getting a red note:
|
Dave, I have no additional info. to add to the RELEASE NOTES (apart from the tables begin external), so feel free to edit the text to a form that you are comfortable with ... Thanks ! |
|
@JS-WRF-SBM Select "develop" instead of "release-v4.0.3", then click "SAVE". |
|
@JS-WRF-SBM
|
I'm following exactly your guidance, but after switching to "develop", I get the above mentioned red note at the top of the screen. The 'SAVE' button disappears (only the EDIT remains) |
Dave, |
|
@JS-WRF-SBM |
|
Hi Dave, I've performed small changes to a single file (reading the initialization look-up tables) and made another 'push' to the newFSBM branch. |
|
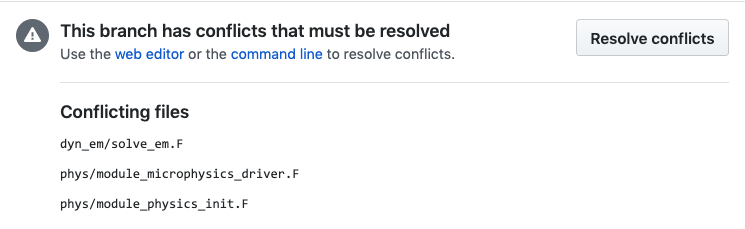
When I try to change to develop it looks like it will allow it but there is a message about changing the base, so I didn't try it. |
Jimy, If you press the button, you will encounter the message that Dave and myself got without any change in the base. |
|
Kobby,
I and Dave looked at your fork again and we see no changes from your
develop to your newFSBM branch,
so it may be that you have not done a git add and git commit on your
changes or have not pushed the new
branch to your fork.
Jimy
…On Wed, Apr 3, 2019 at 10:48 AM JS-WRF-SBM ***@***.***> wrote:
When I try to change to develop it looks like it will allow it but there
is a message about changing the base, so I didn't try it.
Jimy,
If you press the button, you will encounter the message that Dave and
myself got without any change in the base.
—
You are receiving this because you commented.
Reply to this email directly, view it on GitHub
<#848 (comment)>, or mute
the thread
<https://github.com/notifications/unsubscribe-auth/ARGf_CIzuSb8r7WONl_jEUr9kiyHW6o9ks5vdNtwgaJpZM4cVrX1>
.
|
Jimy / Dave, Thanks for pointing this out -- now I can see my new code in the 'newFSBM' branch and the base branch is "develop". Please let me know if anything else needs a change. |
|
Thanks the changes are visible now. |
|
It could be tabs and blank spaces in the Makefile
…On Thu, Apr 4, 2019 at 9:49 AM Dave Gill ***@***.***> wrote:
***@***.**** commented on this pull request.
------------------------------
In phys/Makefile
<#848 (comment)>:
> @@ -36,7 +36,7 @@ MODULES = \
module_bl_shinhong.o \
module_bl_mrf.o \
module_bl_gfs.o \
- module_bl_gfsedmf.o \
+ module_bl_gfsedmf.o \
@JS-WRF-SBM <https://github.com/JS-WRF-SBM>
Kobby,
What ever this change is, we do not want it.
—
You are receiving this because you modified the open/close state.
Reply to this email directly, view it on GitHub
<#848 (review)>,
or mute the thread
<https://github.com/notifications/unsubscribe-auth/ARGf_DEnuumuoOqdxMumQRSDkVijQW47ks5vdh8WgaJpZM4cVrX1>
.
|
|
@JS-WRF-SBM In the code module_mp_SBM_polar_radar.F, "usetables" is a integer variable, and it is later used as a logical variable. Below is the piece of code related to the issue: We need to specify "usetables(ispecies) = 0 or 1" Please take a look. Thanks. |
|
Hi:
Can you idenify yourself?
Do you work with Dave and Jimmy.
Thanks,
Kobby
…On Mon, Jan 27, 2020, 18:52 smileMchen ***@***.***> wrote:
@JS-WRF-SBM <https://github.com/JS-WRF-SBM>
Kobby,
I am running regression test and found that all the compiling failed.
Error messages are as follows:
module_mp_SBM_polar_radar.f90:177:24:
if((ispecies==1) .AND. usetables(ispecies)) then
1
Error: Operands of logical operator '.and.' at (1) are LOGICAL(4)/INTEGER(4)
module_mp_SBM_polar_radar.f90:207:26:
elseif(ispecies==2 .AND. usetables(ispecies)) then
1
Error: Operands of logical operator '.and.' at (1) are LOGICAL(4)/INTEGER(4)
module_mp_SBM_polar_radar.f90:239:26:
elseif(ispecies==3 .AND. usetables(ispecies)) then
1
Error: Operands of logical operator '.and.' at (1) are LOGICAL(4)/INTEGER(4)
module_mp_SBM_polar_radar.f90:324:26:
....
In the code module_mp_SBM_polar_radar.F, "usetables" is a integer
variable, and it is later used as a logical variable. Below is the piece of
code related to the issue:
DO ispecies=1,size(usetables)
if((ispecies==1) .AND. usetables(ispecies)) then ! rain
usetables(1)=0
elseif(ispecies==2 .AND. usetables(ispecies)) then ! fd
usetables(2)=0
elseif(ispecies==3 .AND. usetables(ispecies)) then ! ice crystals (plates, dendrites, columns)
usetables(3)=0
usetables(3)=0
usetables(3)=0
elseif(ispecies==4 .AND. usetables(ispecies)) then ! snow (aggregates)
usetables(4)=0
elseif(ispecies==5 .AND. usetables(ispecies)) then ! graupel
usetables(5)=0
elseif(ispecies==6 .AND. usetables(ispecies)) then ! hail
usetables(6)=0
We need to specify "usetables(ispecies) = 0 or 1"
Please take a look. Thanks.
—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
<#848?email_source=notifications&email_token=ALSMPQVYYDWVVNAH4WDMGQ3Q74GN5A5CNFSM4HCWWX22YY3PNVWWK3TUL52HS4DFVREXG43VMVBW63LNMVXHJKTDN5WW2ZLOORPWSZGOEKAHBDY#issuecomment-578842767>,
or unsubscribe
<https://github.com/notifications/unsubscribe-auth/ALSMPQTW5AEXOHQKZXAVZV3Q74GN5ANCNFSM4HCWWX2Q>
.
|
|
@JS-WRF-SBM @davegill Note that this code can be successfully compiled using intel. However, it failed using GNU. |
|
Also, regression failed for all compiling options (serial, openmp and mpi). |
|
Kobby, yes, smileMchen is here at NCAR working with us. |
|
@JS-WRF-SBM @dudhia @weiwangncar @smileMchen You have far too many changes that are not necessary in solve_em, the MP driver, and diags driver. |
@davegill @dudhia @weiwangncar @smileMchen Thanks for pointing this out. |
|
I don’t think we should merge the code if it cannot be compiled. |
|
Indeed, I meant after resolving the GNU compiler (gfortran) issues. |
|
The unnecessary white space changes should also be eliminated because they
affect our file history.
Jimy
…On Tue, Feb 4, 2020 at 9:33 PM Jacob Shpund ***@***.***> wrote:
@smileMchen <https://github.com/smileMchen>
Indeed, I meant after resolving the GNU compiler (gfortran) issues.
I hardly can assist with this as I do not work with gfortran.
—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
<#848?email_source=notifications&email_token=AEIZ77HGHRZMU5T3GWSGC73RBI6QJA5CNFSM4HCWWX22YY3PNVWWK3TUL52HS4DFVREXG43VMVBW63LNMVXHJKTDN5WW2ZLOORPWSZGOEK2DOAI#issuecomment-582235905>,
or unsubscribe
<https://github.com/notifications/unsubscribe-auth/AEIZ77ETC57DBNLFTU4XVU3RBI6QJANCNFSM4HCWWX2Q>
.
|
|
@JS-WRF-SBM @davegill |
|
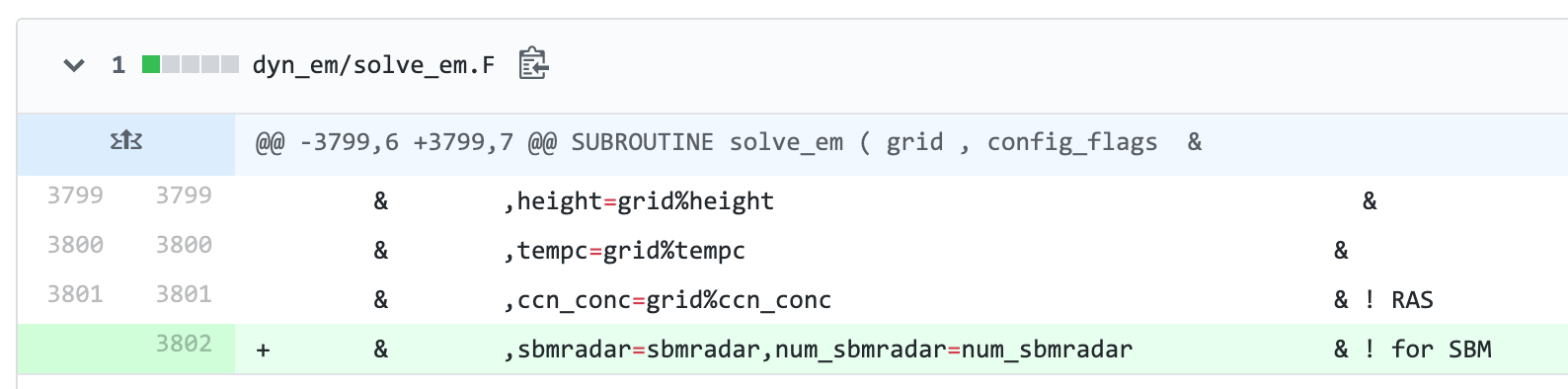
@JS-WRF-SBM Here is the only line to change in solve_em.F: You are changing HUNDREDS of lines with whitespace in several files.
|
|
This removal of white space differences should be a priority for the pull
request.
It seems to apply to all of the common WRF modules that have been changed.
…On Thu, Feb 6, 2020 at 11:25 AM Dave Gill ***@***.***> wrote:
@JS-WRF-SBM <https://github.com/JS-WRF-SBM>
Kobby,
Here is the only line to change in solve_em.F:
[image: Screen Shot 2020-02-06 at 11 18 46 AM]
<https://user-images.githubusercontent.com/12666234/73966074-90545780-48d2-11ea-8bb5-0ed8b4781d6b.png>
You are changing HUNDREDS of lines with whitespace in several files.
1. Use the github "Files changed" page to look at each file that is
modified. You can use the gear-looking icon to ignore white space changes.
[image: Screen Shot 2020-02-06 at 11 21 24 AM]
<https://user-images.githubusercontent.com/12666234/73966266-ea551d00-48d2-11ea-8dc4-695776834835.png>
1. Pull down the latest version of your code that Ming just committed.
Use the previous versions of the the files that you are going to adjust for
white space and re-insert the single line of changes. Your diffs compared
to the version of the develop branch at the time you cut over the mods
should show only the purposeful mods that you intend - not white space
changes.
—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
<#848?email_source=notifications&email_token=AEIZ77FIZG3BL2FX5S2PXPLRBRIX3A5CNFSM4HCWWX22YY3PNVWWK3TUL52HS4DFVREXG43VMVBW63LNMVXHJKTDN5WW2ZLOORPWSZGOELAIPJI#issuecomment-583042981>,
or unsubscribe
<https://github.com/notifications/unsubscribe-auth/AEIZ77HNYUIXR44C7YGL2ZDRBRIX3ANCNFSM4HCWWX2Q>
.
|
|
Jenkin's test yielded multiple failures related to both ideal (em_quarter_ss) and real (double precision) cases. These failures are related to failed compilation. |
|
@smileMchen |
|
newFSBM cannot be compiled for ideal case em_quarter_ss using gnu (as shown in Jenkins test and a separate compiling I did in cheyenne). However, there is no problem when compiled with intel. |
|
The newSFBM codes can be successfully compiled using GNU-v9.1.0. Note that older version of GNU compiler doesn't work due to some internal compiler errors. |
|
@JS-WRF-SBM Unfortunately we just come up with a new problem: regression test failed because the code "module_mp_SBM_polar_radar.F" cannot be compiled with double precision. We need you to fix this issue by the end of this week (Feb. 14), when is the deadline for us to freeze the codes for release. It is fine if you cannot make it by Feb. 14. We can incorporate the newFSBM in the future when it ready. Let us know what you think. |
|
Folks, thanks for your efforts so far in merging the updated FSBM scheme, and apologies for the relatively slow reply as I'm moving to the US on the 14th February for my new position. There is a very little chance that I will be able to do it by the 14th Feb. What is the timescale for resubmit the new PR? Should I wait for the next major release? |
|
@JS-WRF-SBM @smileMchen @dudhia
|
|
I am assuming the polar radar part is not critical to the new SBM.
Is there a way to deactivate it and remove that part of the code?
Jimy
…On Mon, Feb 10, 2020 at 6:22 PM Dave Gill ***@***.***> wrote:
@JS-WRF-SBM <https://github.com/JS-WRF-SBM> @smileMchen
<https://github.com/smileMchen> @dudhia <https://github.com/dudhia>
Kobby,
1. No need to withdraw anything, we'll just keep the PR for the
develop branch.
2. You should look at the PR that Ming put together as a replacement
(getting rid of the extra white space).
3. To replicate the error, just add make the configure step ./configure
-r8
—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
<#848?email_source=notifications&email_token=AEIZ77EP5HWQKNPASCZDZHLRCH4WZA5CNFSM4HCWWX22YY3PNVWWK3TUL52HS4DFVREXG43VMVBW63LNMVXHJKTDN5WW2ZLOORPWSZGOELK5NCI#issuecomment-584439433>,
or unsubscribe
<https://github.com/notifications/unsubscribe-auth/AEIZ77ARLVVGRYZXF6FFKELRCH4WZANCNFSM4HCWWX2Q>
.
|
…1097) TYPE: enhancement / new features KEYWORDS: cloud microphysics, polarimetric-forward-operator SOURCE: Jacob Shpund, Alexander Khain, and Barry Lynn (The Hebrew University of Jerusalem) DESCRIPTION OF CHANGES: A new updated Fast Spectral Bin Microphysics (FSBM) cloud microphysical scheme. There are updates across all the FSBM microphysical scheme including: 1. switch to use either graupel or hail, condensation/evaporation, nucleation (cloud base nucleation, 3 log-normal user-defined aerosol distribution), 2. adaptive cond./evap. time-step, 3. updated collision-coalescence, 4. spontaneous rain breakup, 5. spontaneous snow breakup. A forward polarimetric operator is coupled to the FSBM scheme. The user can see the total reflectivity field, as well as the per hydrometeor total reflectivity (rain, snow, graupel/hail). Please see mandatory input tables info in the RELEASE NOTE. Modified PR from the original #848 and then the additionally modified PR #1085 A new version of the FAST (33 bins) Spectral-Bin Microphysics (FSBM). From #848 -> #1085 1. Inadvertent white space removed 2. Build with NMM 3. Remove -r8 option From #1085 to #1097 1. Fix conflict introduced with #1095 "Separating new_diagnostics from the rest in diag_misc module" LIST OF MODIFIED FILES: M Makefile M Registry/registry.em_shared_collection A Registry/registry.polrad M Registry/registry.sbm M arch/postamble M configure M dyn_em/solve_em.F M phys/Makefile M phys/module_diag_nwp.F M phys/module_diagnostics_driver.F M phys/module_microphysics_driver.F A phys/module_mp_SBM_polar_radar.F M phys/module_mp_fast_sbm.F M phys/module_physics_init.F M share/module_check_a_mundo.F TESTS CONDUCTED: 1. Automated jenkins regression test is all PASS. 2. The code has been compiled and used successfully for 10h of deep convection of the 20th May 2011 MC3E campaign test case (a manuscript consist of scheme details has been submitted for publication). * Compiled using Intel compiler 2019 (ifort version 19.0.3.199) * Configured using options 15 INTEL (ifort/icc, dmpar) and nest option 1. * BC/IC used the NCEP FNL (Final) Operational Global Analysis data are on 1x1 degree grids - Domain setup and time step (5s per 1.3 km): ``` &domains time_step = 15, time_step_fract_num = 0, time_step_fract_den = 1, max_dom = 2, dx = 4000, 1333.33, dy = 4000, 1333.33, grid_id = 1, 2, parent_id = 0, 1, parent_grid_ratio = 1, 3, parent_time_step_ratio = 1, 3, ``` - In order to use the bin-wise polarimetric forward operator, a new flag must exist in the <namelist.input> file: ``` &physics sbm_diagnostics = 1, 1, ``` - The FAST SBM code is not currently configure to work with default 8-byte reals. RELEASE NOTE: Fast (33 bins) Spectral-bin Microphysics (FSBM): in order to run the new FSBM scheme, users needs to download an external directory named "SBM_input_33" consist of mandatory input tables and place it in the 'run' directory. In case the coupled polarimetric forward operator is to be used (e.g., 'sbm_diagnostics = 1'), a second directory consist of scattering amplitudes named "SBM_scatter_amplit.tgz" needs to be placed in the 'run' directory. Both directories are compressed and can be downloaded at the following link: https://drive.google.com/drive/folders/1qxYyQwKI1wKQYasDUkQvVgHs11prLiqA?usp=sharing (on a Linux machine, you may need to escape the `?` character with `\?`). The FAST SBM code is not currently configure to work with default 8-byte reals.
|
Replaced with #1097 |

A new version of the FAST (33 bins) Spectral-Bin Microphysics (FSBM)
TYPE: enhancement / new features
KEYWORDS: cloud microphysics, polarimetric-forward-operator
SOURCE: Jacob Shpund, Alexander Khain and Barry Lynn (The Hebrew University of Jerusalem)
DESCRIPTION OF CHANGES: A new updated FSBM cloud microphysical scheme.
There are updates across all the FSBM microphysical scheme including: switch to use either graupel or hail, condensation/evaporation, nucleation (cloud base nucleation, 3 log-normal user-defined aerosol distribution), adaptive cond./evap. time-step, updated collision-coalescence, spontaneous rain breakup, spontaneous snow breakup.
A forward polarimetric operator is coupled to the FSBM scheme, where the user can see the total reflectivity field, as well as the per hydrometeor total reflectivity (rain, snow, graupel/hail). Please see mandatory input tables info in the RELEASE NOTE.
ISSUE:
LIST OF MODIFIED FILES:
M Registry/registry.em_shared_collection
A Registry/registry.polrad
M Registry/registry.sbm
M dyn_em/solve_em.F
M phys/Makefile
M phys/module_diag_misc.F
M phys/module_diagnostics_driver.F
M phys/module_microphysics_driver.F
A phys/module_mp_SBM_polar_radar.F
M phys/module_mp_fast_sbm.F
M phys/module_physics_init.F
TESTS CONDUCTED: The WTF has not been used yet for the updated FSBM code.
However, the code has been compiled and used successfully for 10h of deep convection of the 20th May 2011 MC3E campaign test case (a manuscript consist of scheme details has been submitted for publication):
RELEASE NOTE:
Fast (33bins) Spectral-bin Microphysics (FSBM): in order to run the new FSBM scheme, users needs to download an external directory named "SBM_input_33" consist of mandatory input tables and place it in the 'run' directory.
In case the coupled polarimetric forward operator is to be used (e.g., 'sbm_diagnostics = 1'), a second directory consist of scattering amplitudes named "SBM_scatter_amplit.tgz" needs to be placed in the 'run' directory.
Both directories are compressed and can be downloaded at the following link:
https://drive.google.com/drive/folders/1qxYyQwKI1wKQYasDUkQvVgHs11prLiqA?usp=sharing