-
-
Notifications
You must be signed in to change notification settings - Fork 1.5k
BUG: Fix bug with scaling and fids for standard montages #10324
New issue
Have a question about this project? Sign up for a free GitHub account to open an issue and contact its maintainers and the community.
By clicking “Sign up for GitHub”, you agree to our terms of service and privacy statement. We’ll occasionally send you account related emails.
Already on GitHub? Sign in to your account
Changes from all commits
File filter
Filter by extension
Conversations
Jump to
Diff view
Diff view
There are no files selected for viewing
| Original file line number | Diff line number | Diff line change |
|---|---|---|
|
|
@@ -29,10 +29,11 @@ def test_standard_montages_have_fids(kind): | |
| for k, v in fids.items(): | ||
| assert v is not None, k | ||
| for d in montage.dig: | ||
| if kind == 'artinis-octamon' or kind == 'artinis-brite23': | ||
| assert d['coord_frame'] == FIFF.FIFFV_COORD_MRI | ||
| if kind.startswith(('artinis', 'standard', 'mgh')): | ||
| want = FIFF.FIFFV_COORD_MRI | ||
| else: | ||
| assert d['coord_frame'] == FIFF.FIFFV_COORD_UNKNOWN | ||
| want = FIFF.FIFFV_COORD_UNKNOWN | ||
| assert d['coord_frame'] == want | ||
|
|
||
|
|
||
| def test_standard_montage_errors(): | ||
|
|
@@ -142,12 +143,11 @@ def test_set_montage_artinis_fsaverage(kind): | |
| assert trans['from'] == trans_fs['from'] | ||
| translation = 1000 * np.linalg.norm(trans['trans'][:3, 3] - | ||
| trans_fs['trans'][:3, 3]) | ||
| # TODO: This is actually quite big... | ||
| assert 15 < translation < 18 # mm | ||
| assert 0 < translation < 1 # mm | ||
| rotation = np.rad2deg( | ||
| _angle_between_quats(rot_to_quat(trans['trans'][:3, :3]), | ||
| rot_to_quat(trans_fs['trans'][:3, :3]))) | ||
| assert 3 < rotation < 7 # degrees | ||
| assert 0 < rotation < 1 # degrees | ||
|
|
||
|
|
||
| def test_set_montage_artinis_basic(): | ||
|
|
@@ -172,15 +172,15 @@ def test_set_montage_artinis_basic(): | |
|
|
||
| # Check a known location | ||
| assert_array_almost_equal(raw.info['chs'][0]['loc'][:3], | ||
| [0.0616, 0.075398, 0.07347]) | ||
| [0.054243, 0.081884, 0.054544]) | ||
|
Member
Author
There was a problem hiding this comment. Choose a reason for hiding this commentThe reason will be displayed to describe this comment to others. Learn more. @HanBnrd @rob-luke are the
Contributor
There was a problem hiding this comment. Choose a reason for hiding this commentThe reason will be displayed to describe this comment to others. Learn more. Hi @larsoner, yes artinis-brite23.elc and artinis-octamon.elc are using MNI coordinates. Sorry, what would you like to check exactly? Would you like us to run
Member
Author
There was a problem hiding this comment. Choose a reason for hiding this commentThe reason will be displayed to describe this comment to others. Learn more.
Yeah that would be great. The changes suggest that they will shift anterior (and a little inferior) by ~1.5 cm but it should actually be more correct
Contributor
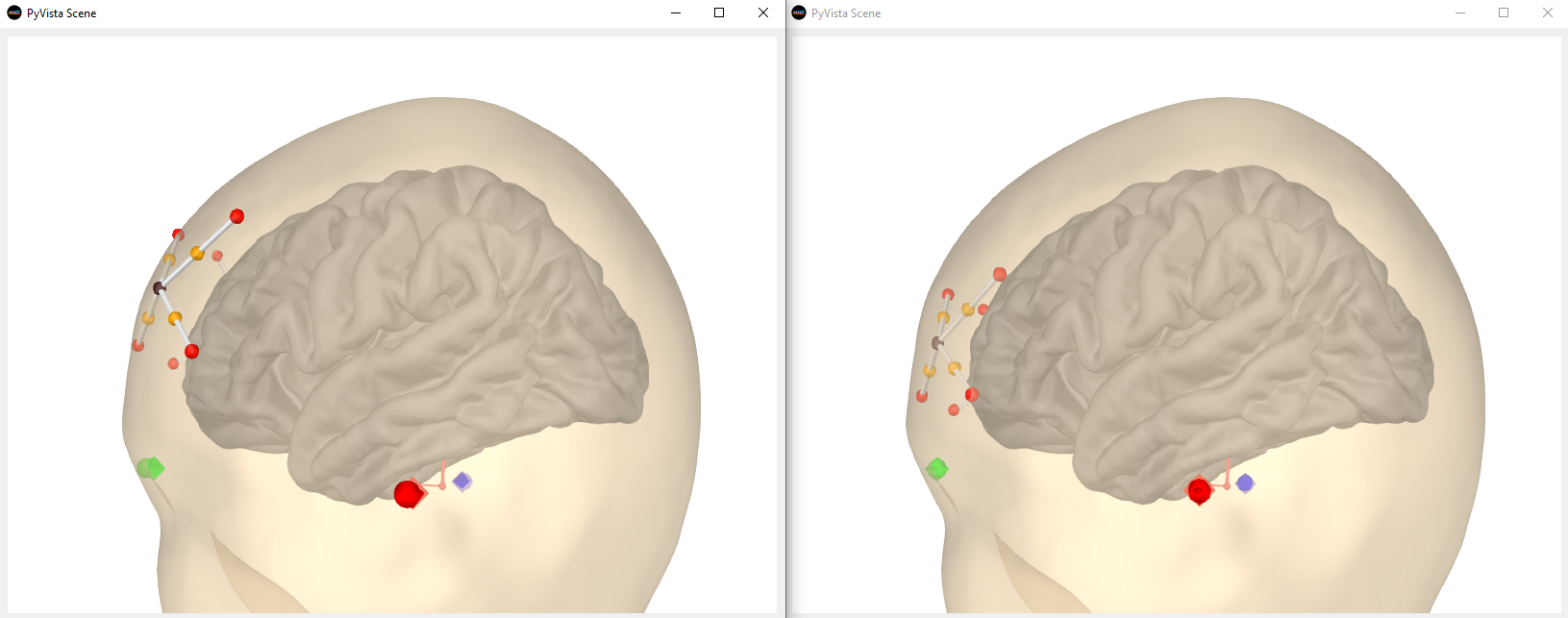
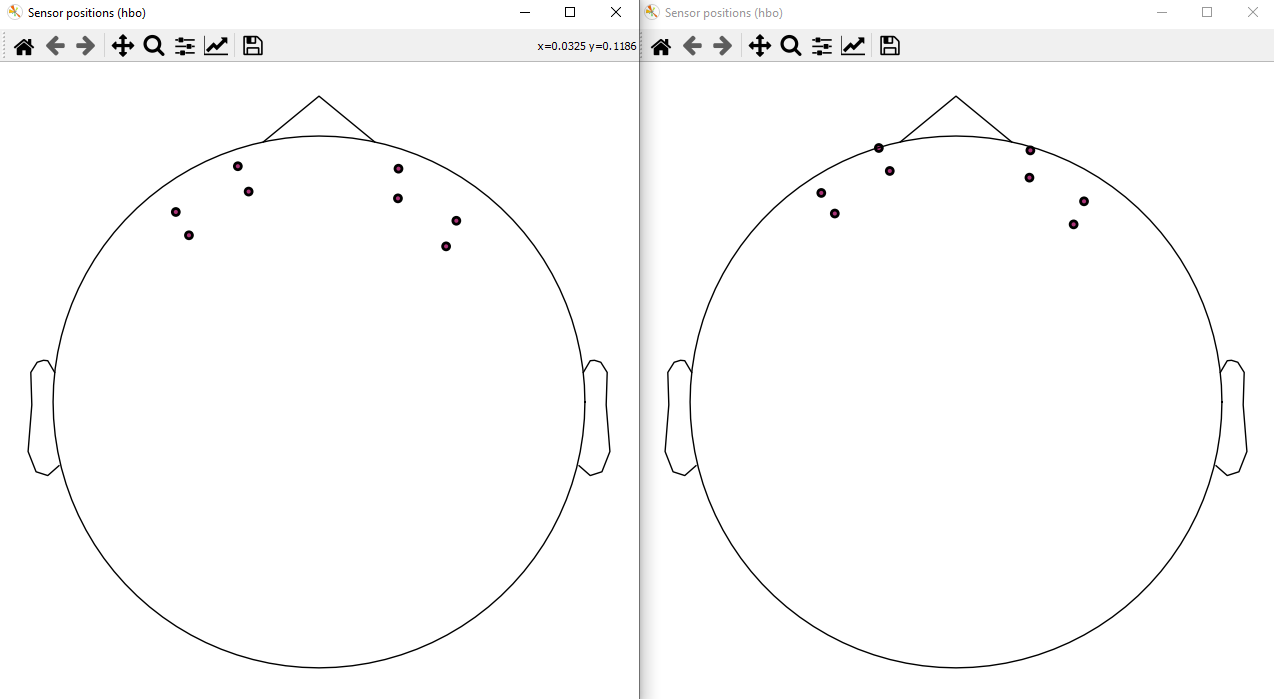
There was a problem hiding this comment. Choose a reason for hiding this commentThe reason will be displayed to describe this comment to others. Learn more. Hi @larsoner, this is before (left) and after (right) the PR with the Artinis Octamon montage It seems that after the PR, the sensors look like they're inside the head in the 3D vis but this is a bit confusing because the 2D viz shows the sensors more anterior indeed. This is the code I used: raw.set_montage('artinis-octamon')
raw.plot_sensors()
subjects_dir = mne.datasets.sample.data_path() + '/subjects'
mne.datasets.fetch_fsaverage(subjects_dir=subjects_dir, verbose=True)
fig = mne.viz.create_3d_figure(size=(800, 600), bgcolor='white')
fig = mne.viz.plot_alignment(raw.info, show_axes=True,
subject='fsaverage', coord_frame='mri',
trans='fsaverage', surfaces=['brain', 'head'],
fnirs=['channels', 'pairs',
'sources', 'detectors'],
dig=True, mri_fiducials=True,
subjects_dir=subjects_dir, fig=fig)
mne.viz.set_3d_view(figure=fig, azimuth=70, elevation=100, distance=0.4,
focalpoint=(0., -0.01, 0.02))
Member
Author
There was a problem hiding this comment. Choose a reason for hiding this commentThe reason will be displayed to describe this comment to others. Learn more.
It could be that the head surface that the Artinis folks used to create the montage was slightly different. This seems to be the case for our EEG standard montages at least. It's tough to know which one is "more correct" I guess... |
||
| assert_array_almost_equal(raw.info['chs'][8]['loc'][:3], | ||
| [-0.033875, 0.101276, 0.077291]) | ||
| [-0.03013, 0.105097, 0.055894]) | ||
| assert_array_almost_equal(raw.info['chs'][12]['loc'][:3], | ||
| [-0.062749, 0.080417, 0.074884]) | ||
| [-0.055681, 0.086566, 0.055858]) | ||
| assert_array_almost_equal(raw_od.info['chs'][12]['loc'][:3], | ||
| [-0.062749, 0.080417, 0.074884]) | ||
| [-0.055681, 0.086566, 0.055858]) | ||
| assert_array_almost_equal(raw_hb.info['chs'][12]['loc'][:3], | ||
| [-0.062749, 0.080417, 0.074884]) | ||
| [-0.055681, 0.086566, 0.055858]) | ||
| # Check that locations are identical for a pair of channels (all elements | ||
| # except the 10th which is the wavelength if not hbo and hbr type) | ||
| assert_array_almost_equal(raw.info['chs'][0]['loc'][:9], | ||
|
|
@@ -200,11 +200,11 @@ def test_set_montage_artinis_basic(): | |
| raw.info['chs'][0]['loc'][:9]) | ||
| # Check a known location | ||
| assert_array_almost_equal(raw.info['chs'][0]['loc'][:3], | ||
| [0.085583, 0.036275, 0.089426]) | ||
| [0.068931, 0.046201, 0.072055]) | ||
| assert_array_almost_equal(raw.info['chs'][8]['loc'][:3], | ||
| [0.069555, 0.078579, 0.069305]) | ||
| [0.055196, 0.082757, 0.052165]) | ||
| assert_array_almost_equal(raw.info['chs'][12]['loc'][:3], | ||
| [0.044861, 0.100952, 0.065175]) | ||
| [0.033592, 0.102607, 0.047423]) | ||
| # Check that locations are identical for a pair of channels (all elements | ||
| # except the 10th which is the wavelength if not hbo and hbr type) | ||
| assert_array_almost_equal(raw.info['chs'][0]['loc'][:9], | ||
|
|
||
There was a problem hiding this comment.
Choose a reason for hiding this comment
The reason will be displayed to describe this comment to others. Learn more.
This is a sign that things work better (and that they were broken before by this amount)