-
Notifications
You must be signed in to change notification settings - Fork 14
New issue
Have a question about this project? Sign up for a free GitHub account to open an issue and contact its maintainers and the community.
By clicking “Sign up for GitHub”, you agree to our terms of service and privacy statement. We’ll occasionally send you account related emails.
Already on GitHub? Sign in to your account
Question on the meaning of parameters of generalised_geodesic3d #40
Comments
|
Hi @anafyp , Thats a great question, many thanks for raising it. This library is mainly aimed at calculating Geodesic distance transforms with a fast parallelized implementation. There is a trade-off between accuracy and speed, where most fast implementations are an approximation for the actual Geodesic distance transform. This also applies if you are doing Euclidean distance with the functions here, they may be fast but not exact computation but rather a close approximation. Having said that, the library can be used to compute Euclidean distances. You need to provide an empty image (doesnt not matter what values you provide as for Euclidean distance this is not used but still required by the functions) and setting lamb = 0.0 will enable computing the Euclidean distance transform, which is similar to scipy edt function. Here is an example I implemented for your reference FastGeodisEuclideanDistanceTransformWithoutImage Regarding your question about other parameters, please have a look at the simple demo colab samples provided here: Colab samples I provide a brief overview about each parameter for your Euclidean distance use case:
I hope this helps. |
|
I forgot to add, if there is an interest in having specific Euclidean distance only functions, I can implement and include those within the library as well (so you wont need to set all the parameters that are specifically aimed at Geodesic distance) |
|
Thank you @masadcv for your response, that is also the behavior that I expected. The result however is very different from the one by scipy. FastGeodis with the parameters you recommended produces the following output: While scipy, with the expected result produces: I think there is indeed an interest in having a specific Euclidian distance-only function, but for now, this distance looks more like a chessboard distance and not a euclidian one. |
|
Can you share this example so I can have a closer look at what you are trying? |
|
As i am working with medical data it might be hard to share data... I was mistaken in my previous comment I think FastGeodis computes the taxicab/manhattan distance and not the euclidian one. Here is a dummy example: |
|
As discussed earlier, on the long run, it would be good to have dedicated approaches for Euclidean distance maps. Tensorflow has a GPU implementation of the Felzenszwalb & Huttenlocher Distance transform which could be ported here:
CUCIM has a GPU implementation of the Parallel Banding Algorithm (PBA) which may be even more appropriate for porting here: By the way, is the current output from Fast Marching any better? |
|
Many thanks @tvercaut for pointing out the reference implementations we can use for Euclidean distance maps. I have not checked Fast Marching for Euclidean distance maps yet. Will check and see if it is any better. |
|
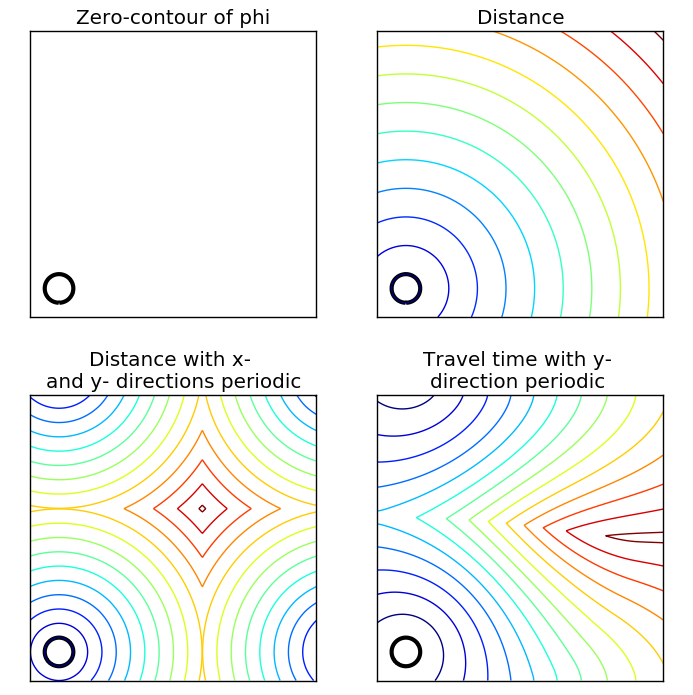
If it's of help for comparison purposes, the scikit-fmm package provide Euclidean distance map computation through fast marching: As far as their demo shows, their implementation doesn't seem to lead to star artefacts: |
|
Hi @anafyp , Could you try your examples with version 1.0.3 (make sure to refer to API docs to correct any function naming/arguments mismatch in new version) |






Hi all,
I have recently discovered your implementation of FastGeodis. I am however unable to understand how I am supposed to make it work.
Say we have a 3D binary mask in tensor format (i.e [B, C, X, Y, Z]) if I wanted to simply do the distance transform of this mask as in scipy.ndimage.morphology.distance_transform_edt, how should I proceed?
To go further with my question I don't really understand the documentation on:
To what should I set those if I just want to extract the euclidian distance transform of my mask ?
Thanks!
The text was updated successfully, but these errors were encountered: