Explicit 2D Diffusion Correlation Operator#966
Conversation
|
@travissluka, Awesome, great job. This algorithm is not trivial. Congratulations! The good thing is that this pseudo-diffusion operator is self-adjoint and passes the self_adjoint tests. I am glad that ROMS algorithms were useful. Implementing the methodologies we see in papers into discrete models is usually complicated and takes time. Yes, the randomization approach is the way to go. I recall it took me about two days to compute those coefficients in a large grid using the exact method. By the way, I have checked and used the coding in SOCA that you and others have developed many times as a prototype in the ROMS-JEDI interface. |
| end subroutine | ||
|
|
||
| ! ------------------------------------------------------------------------------ | ||
| ! Calibration for the explicit diffuion operator. |
There was a problem hiding this comment.
ppppffffff .... difffuzzion not diffuion 🤦♂️
 guillaumevernieres
left a comment
guillaumevernieres
left a comment
There was a problem hiding this comment.
🎉
Thanks for doing this @travissluka ! Looks good as always. If there are bugs we'll find them by putting that chunk of code through its paces.
Preemptively approving before the CI finishes.
PS: Did you check within one of the variational application too @travissluka ?
|
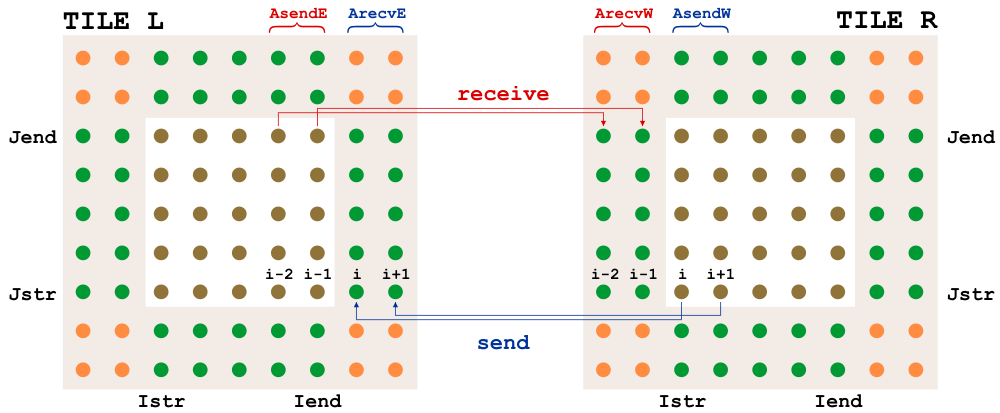
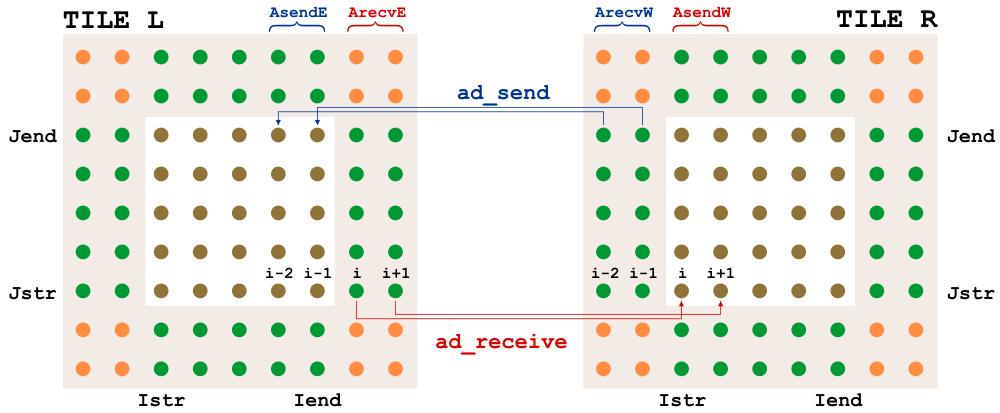
@travissluka, looking for parallel bugs is time-consuming; good luck. Hopefully, it will be easy to find them by putting some print statements about the i- and j-ranges. Otherwise, it is looking good. Knowing that the explicit diffusion operator is faster than BUMP is useful. In ROMS, we can have either 2 or 3 halo points. The diagrams for NLM/TLM and ADM for 2 halo points are: |
|
@travissluka, Great. Oh yeah, I remember generating random numbers in parallel is very tricky. I struggled a lot with getting an identical solution with different parallel partitions. We have a very convoluted way of creating random numbers for our algorithms. |





Description
(well that took longer than expected!, and not at all because I accidentally deleted half the code last week)
This PR creates a
SaberCentralBlockfor a soca-specific 2D explicit diffusion operator. (Following Weaver 2001, and pulling inspiration from the ROMS code). It is meant to handle the short correlation lengths. For longer lengths, it's better to use BUMPMuch like BUMP, using this block is a two-step process
test/testinput/parameters_diffusion.yml). Two ways of doing normalization are possible.brute forcecalculates the normalization exactly, but as you might guess from the name, you don't want to do this for a grid bigger than the 5 deg ctest grid. It's just there to check the code.randomizationrandom vectors are generated, and the TL of the diffusion operator is applied, to estimate the normalization values. (~10,000 iterations is good for a real case)test/testinput/dirac_diffusion.yml) the previous parameter file is read in.other changes that were made to get this to work
setcorscaleapplication (how the rossby radius based length were specified) was modified to mirror exactly what is used insoca_covariance_mod. Required a small change in thesetcorscale.yaml(min->min value)gridgenwill now also savedxanddyso that they can be used in this filter.existing soca_gridspec.nc files will need to be updated after the PR
Testing
The following tests were done on my own machine
ctest
2 ctests are added, one for calibration, one for a dirac test.
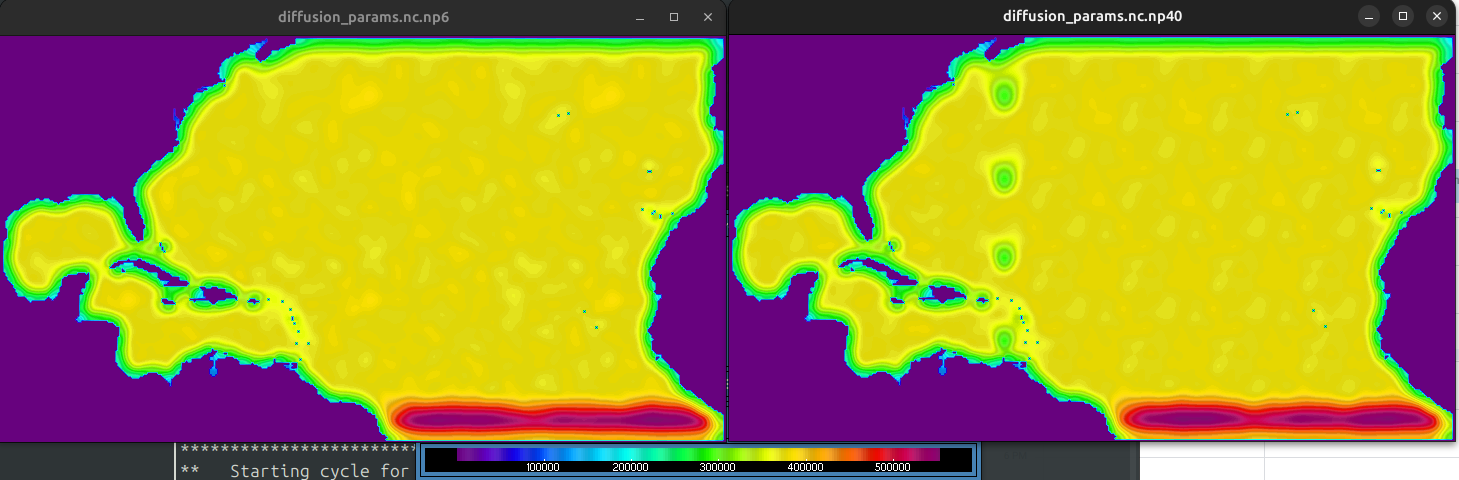
brute force, you'll see in the dirac test that everything is perfectly normalized. The value at the dirac location stays 1.0 🎉src/soca/Fields/soca_fields_mod.F90you'll see that the self-adjoint test does pass 🎉 . The adjoint test can't be left on because it fails if the halos are modified. (soca'stoFieldset()does a halo exchange, and other tests break if I remove that halo exchange). To be fixed another day.realistic scales on a 1 deg grid
absurdly large scales on a 1 deg grid
multiply()!) so don't do this in the real world. But it works, and normalization is still good.extra sanity check
create a dirac with a length scale of 4 gridboxes, and 4 grid boxes away the value is ~0.6 as expected for 1 sigma of a gaussian

1/4 deg grid regional 3dvar
Followup steps
likely future follow-ups to this PRs
This will have the normalization done separately so that 2D normalization can be computed offline (it's slower), and the vertical normalization can be computed each cycle to respond to varying mixed layer depths and such
HorizFiltSOCAtagging other people who were interested: @hga007 @shlyaeva @ncrossette